Why, What, How, Who, What else?
Why look at genetic background by participating in a yDNA study?
We have all experienced hitting the proverbial brick wall in our genealogical research. Those of us with the last name "Cathcart/Kathcart/Kithcart" are no exception. Also, given the nature of Scots-Irish research, only the most lucky of us can trace back with any degree of certainty into the 18th Century. The genealogical records are pretty scarce back in Scotland, England, and especially in Ulster (Northern Ireland).
What if you've only been able to trace back three or four generations? What to do? There often simply aren't enough records to fill in the blanks for those ancestors... and the brick wall looms large and impassable.
Genetic testing gives us another option. Once you find out which male Cathcart's yDNA most closely match your own, you can concentrate your genealogical research in those families most likely related to you in a reasonable number of generations.
The Family Tree DNA website is sometimes unintelligible. Markers, haplo-groups, I mean, do I really care if my marker number 459b is a 9? And what does that mean? J
What I'm going to try to do is allow us to see the results of our tests, and have a look at the genealogy of those most closely related to us genetically speaking.
 Raymond
Cathcart originally set up our Cathcart yDNA group with the company Family Tree DNA
(FTDNA) and still serves as the group co-administrator.
Raymond
Cathcart originally set up our Cathcart yDNA group with the company Family Tree DNA
(FTDNA) and still serves as the group co-administrator.
Rather than try and explain the process myself, please have a look at their website, you'll find most of your questions answered (What it is, how it works, how much it costs, etc.).
- FTDNA Website's FAQ page...
- FTDNA Website's yDNA page...
- Our Cathcart Project Website (public page, no password required)...
- Join the "Cathcart" Group Project...
Other Links
- Excellent Wikipedia article on genealogical DNA testing
- Blair DNA Project: excellent basic background on...
- List of y STR markers (details, mutation rates, sortable list)
A few more points and links:
- None of us have any stock in the company. We support this program to expand our knowledge about our Cathcart heritage... not expand our pocketbooks!
- If you need help with the cost, please let me know, we have collected funds to help defray—or completely cover—the costs of the test.
- If you would like to help us by making a contribution to our group, you may do so on the above website.
- Other genetic testing companies
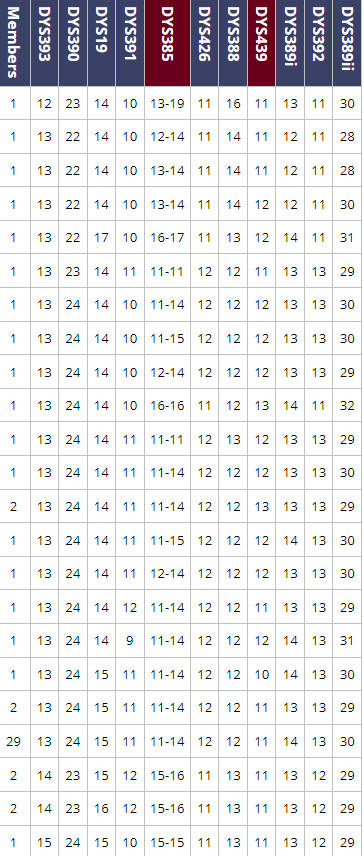 |
| Unique Haplotypes @ 12-marker level as of May 18, 2024 |
- The number of participants continues to grow.
As of today (5/18/2024) our Cathcart study has...
- 74 total participants
- 7 Big Y
- 18 111-marker level
- 42 67-marker level
- 54 37-marker level
- 8 yDNA confirmed haplogroups
- 51 participants have shared their paternal Ancestor information, while 3 have uploaded their combined GEDCOM databases submitted (that is, participants' digital ancestor listings/information)
- 16 distinct yDNA predicted haplogroups:
- 3 have been confirmed by further/more detailed testing of long-lasting SNPs1
- Most of our participants come from the main Haplogroups, R and I.
- We now have two members from the E Group... welcome!
- FamilyTreeDNA has
recently started defining participants' Haplogroup via SNP
designation--either "confirmed" or "predicted."
- So, where this column used to say
Haplogroup R1b1a2, it will now indicate
R-M269.
- Of this group, two of us (Raymond and I) have tested/confirmed R-DF27
- So, where this column used to say
Haplogroup R1b1a2, it will now indicate
R-M269.
- Of the I2 group, two have tested/confirmed I-M223
- Click here for a more detailed discussion
- We've also had 18 mtDNA Haplogroups identified amongst our 21 members whose mtDNA was tested. 30 of our participants have completed a Family Finder (autosomal DNA) test.
- At the outset, our Cathcart group has concentrated on yDNA and therefore on genes passed directly from father to son through the generations. We do this for genetic reasons, not because we're "sexist" or a "males only club!" :-)
- Although mtDNA is passed from a mother to her son; the son will not, in turn pass this genetic material on to his son.
- The study should help some participants and to give more "depth" to our study
- 74 total participants
- What does it mean to match exactly at the
67-marker test level?
- According to FamilyTreeDNA's website, matching at this level (with the same or derivative surname: i.e. Cathcart/Cithcart/Kathcart) indicates that these gentlemen are "very tightly related." In fact, at this level, 50% find a common ancestor within three generations; and that there is a 90% probability that they share a common ancestor within a "mere" five generations!
- Click here for pop-up chart
- Our group also includes several
combinations of exact matches at the 37-, 25-, and
12-marker level. What do these results mean?
- Again, assuming the same/derivative
surname, an exact matches have the following implications:
- at the 37-marker level: a "very tight relation," a 50% possibility of a common ancestor within five generations, and a 95% probability that that ancestor could be found within eight generations.
- at the 25-marker level: participants are "related," and that they "likely share a common ancestor within the 'genealogical timeframe.'"
- Again, assuming the same/derivative
surname, an exact matches have the following implications:
- For more details about how to interpret yDNA test results, please visit the FamilyTreeDNA website's FAQs page.
- This is a flexible page -- which should work in most browsers -- that allows you to select various yDNA results pages in two separate "frames"
- Information available on this page:
- Main Cathcart Groups
- The "Brick Wall" individuals where our genealogical research stops
- yDNA results grouping charts (simple chart showing ancestors back as far as is known via genealogical sources).
- Participants' Pedigree Charts (from external RootsWeb website) if available.
-
yDNA relationship matrix
- A simple chart showing genetic distance results amongst the above "Cathcart Groups"
- Main Cathcart Groups
- All these pages are "under construction," and I'd love to have your feedback!
- use this "stripped down" version if your browser doesn't support frames (which are used on the above-mentioned page, and which may not work in all web browsers).
- Since the company no longer hosts free online
trees, I'm in the process of removing RootsWeb pedigree charts
- Eventually, I hope to link to Ancestry.com trees.
- Please let me know if there are errors in my data! I'll correct them as soon as I can.
Ongoing and Future Enhancements
-
Cathcart DNA Blog
- Ask questions, make comments, share your genetic genealogy discoveries
- Use my contact page or email if you'd rather share in private.
- I have recently added Google map overlays showing which
Cathcarts were where and when to several sections of this website.
- Although I've only added a few locations to these maps (for South Carolina and Tennessee), I hope to add more of these geographical listings over the next several months.
- I've also used Google Maps to identify various locations in County Antrim, Ireland to help Cathcart researchers.
- This is what I was trying to accomplish with my timeline page on this site.
- Now that I've regained my administrative access to the Cathcart yDNA website, I hope to create an online database to permit participants to view and sort results.
- SNP Single-Nucleotide Polymorphism (pronounced
"snip") is a long-lasting DNA sequence that is increasingly being used
to track genetic and population changes. In the past, the genetic
genealogical community used a "phylogenetic" or family
tree based naming system to define Haplogroups. Now it is more common
to use the SNP marker designation. However, one drawback of this newer
system is that it's harder to easily recognize subclades (i.e. related Haplogroups that
begin with the same starting sequence of numbers and letters).
- R1b1a2
is thus replaced by R-M269.
- The included sub clade R1b1a2a1a2 is now typically defined by the mutation SNP marker R-P312 (also known as S116).
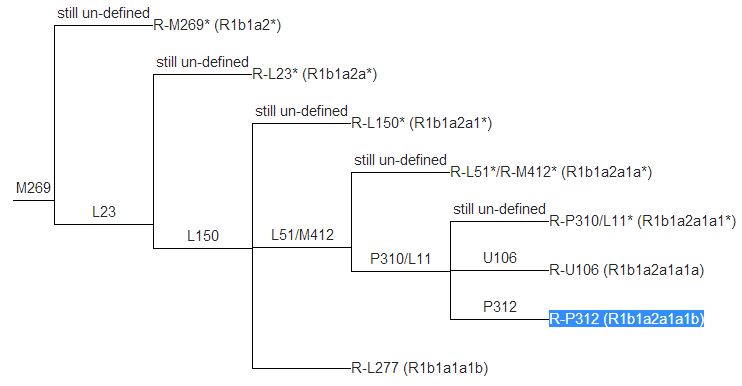
- In the chart below, R-M269 (R1b1a2) is
located on the far left, with specific subclades (branches)
spreading to the right. The most common of our participants'
predicted Haplogroup, R-P312 (R1b1a2a1a1b) is highlighted in blue at
the bottom right.
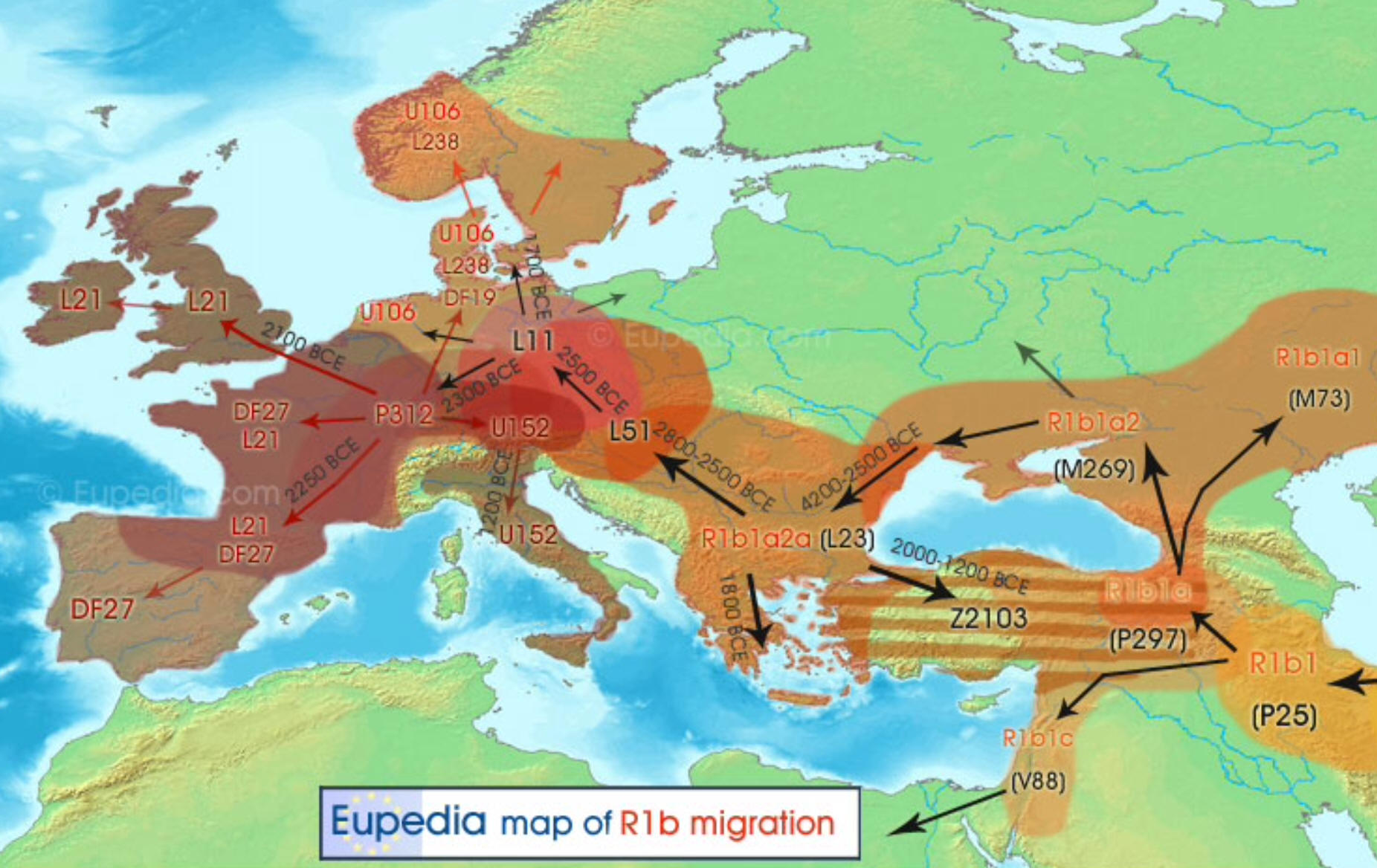
- In the map below, you can see the migration/mutation geographically with notes indicating the timeframe of the mutation(s).
Look for R-M269 just above the Black Sea, and the later R-P312
mutation is just north of the Alps. Note that we are looking at genetic changes from thousands of years ago... not particularly helpful with our genealogical research!
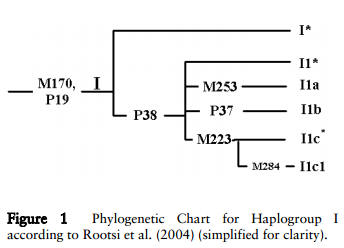
-
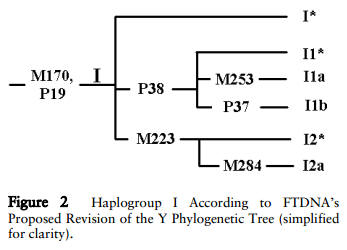
For the participants of Haplogroup I, things get a
bit difficult. There is apparently ongoing debate on just how
to place the two SNP subgroups I-M253 (I1a) and
I-M223 (either I1c or I2*). Here are two charts depicting the
different trees... I'll leave it to the experts to figure out
which one is more correct!:
- R1b1a2
is thus replaced by R-M269.
- By studying past results of specific SNP
markers, FamilyTreeDNA.com can predict the participant's Haplogroup
by studying specific STR (Short Tandem Repeat)
markers. It's not as precise as testing the individual SNPs, but can
give fairly accurate results.
- You can explore your own Haplotree by clicking on the icon on your main welcome page that FamilyTreeDNA.com. from this page you can also order individual SNP tests (currently $39 each).
